| Kinase | Description |
|---|---|
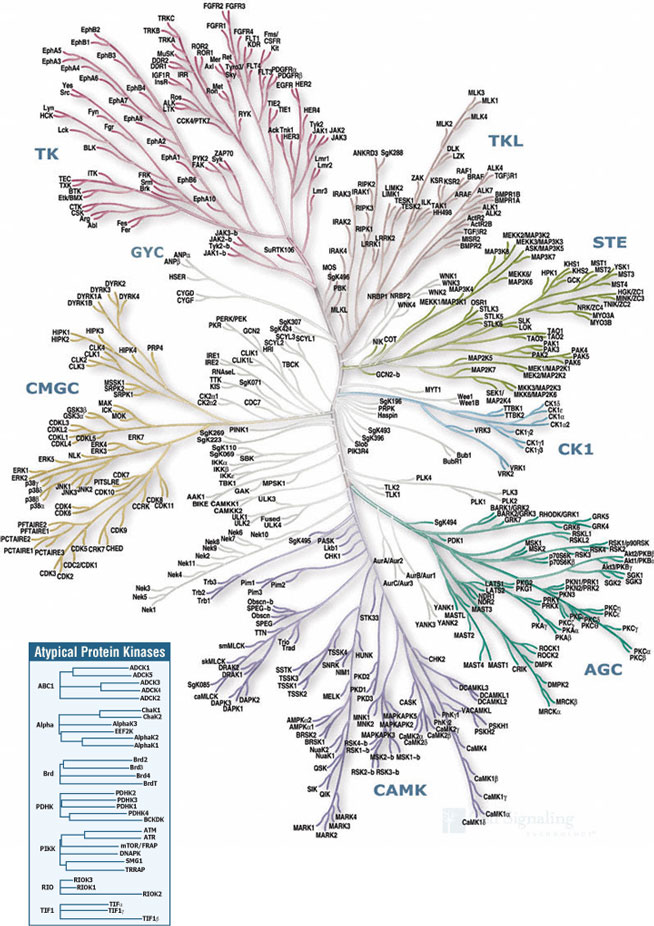
| AGC | Containing PKA, PKG, PKC families |
| CAMK | Calcium/calmodulin-dependent protein kinase |
| CK1 | Casein kinase 1 |
| CMGC | Containing CDK, MAPK, GSK3, CLK families |
| STE | Homologs of yeast Sterile 7, Sterile 11, Sterile 20 kinases |
| TK | Tyrosine kinase |
| TKL | Tyrosine kinase-like |
Click on the individual groups to view the detail page of that group.
To request the use of The Human Kinome by Cell Signaling Technology (CST), please complete our request form.
The main dendrogram shows the sequence similarity between protein kinase domains, derived from public sequences and gene prediction methods detailed in Manning et al. (Science, 298, 1912-1934). Domains were defined by hidden Markov model profile analysis and multiple sequence alignment. The initial branching pattern was built from a neighbor-joining tree derived from a clustalW protein sequence alignment of the domains. This was extensively modified by reference to other alignment and tree building methods (hmmalign and parsimony trees), and extensive pairwise sequence alignment of kinase domains. The curved layout was created manually. Many branch lengths are semi-quantitative, but the branching pattern is more informative than any single automatic method. The more detailed trees were generated automatically by clustalW alignment of full length protein sequences followed by neighbor-joining tree building. Unpublished kinases are named where possible according to family nomenclature. Some divergent kinases retain a numerical SgK (SuGen Kinase) accession number. The second domains of dual-domain kinases are named with a "~b" suffix.